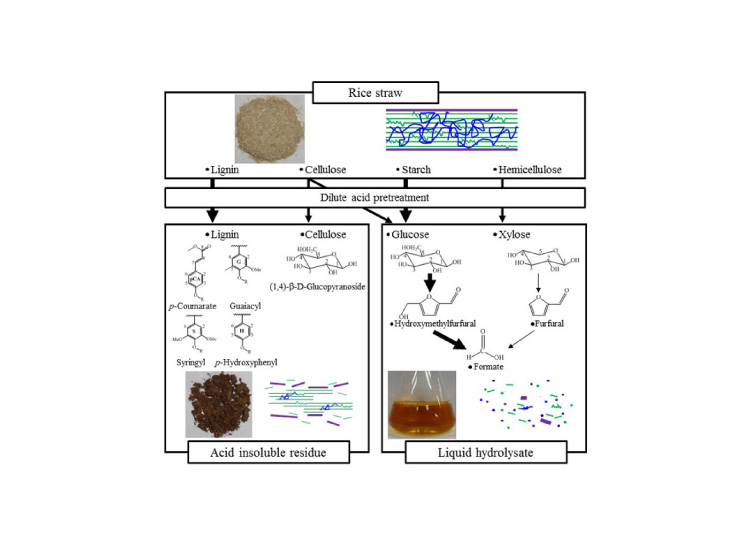
A Japanese research team have applied a method used in human genetic analysis to rice and rapidly discovered four new genes that are potentially significant for agriculture. These findings could influence crop breeding and help combat food shortages caused by a growing population. The paper was published on June 21, 2016 (Japan Standard Time) in the online edition of Nature Genetics.
Selective crop improvement based on plant genetics and breeding is essential to support the world’s growing population. In order to efficiently breed new crop varieties it is necessary to rapidly identify the genes related to high crop yields and analyze what makes them special.
Until now the genetic analysis of crops has mainly been based on quantitative trait loci (QTL) analysis, but this method requires time to develop experimental populations. Another method known as genome-wide association studies (GWAS), frequently used to analyze human genes, uses data from many extant individuals to analyze genes in a short time span. Various plant species have also been analyzed using this method, but there have been very few cases of successful analysis.
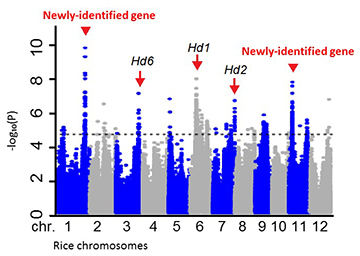
The known genes Hd1, Hd2, and Hd6 were located, together with two newly-identified genes that also affect flowering dates.
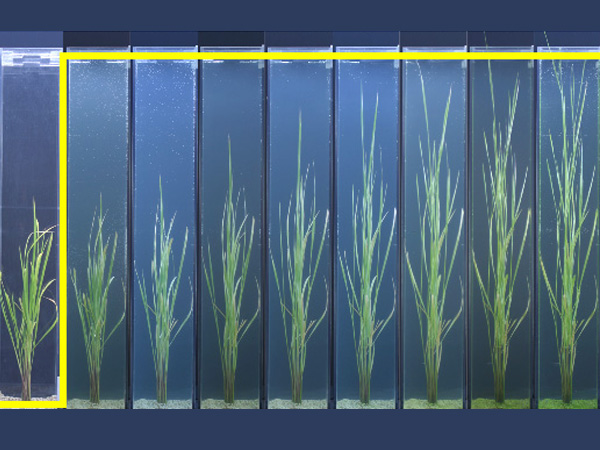
In order to produce results using GWAS analysis, the research team limited their targets to 176 Japanese rice cultivars, including 86 cultivars used in Japanese-sake brewing that Kobe University has maintained over many years. Using next-generation sequencing, the group determined the whole sequence of each cultivar, and discovered a total of 493,881 of the DNA-based polymorphisms.
Based on these results, the team carried out GWAS analysis on each trait and rapidly identified four genes within a group of 12 rice plant chromosomes. Chromosome 1 contains a gene that decides rice flowering date; chromosome 4 contains a gene that influences panicle number produced, leaf breadth, and rice grain number; a chromosome 8 gene affects awn length (a factor which influences harvesting); and a gene within chromosome 11 decides flowering date, plant height, and panicle length.
Genetic analysis of plants based on GWAS has been carried out many times but with limited success. The success of this experiment could aid the discovery of genes in other plant and animal species and potentially contribute towards solving food shortages caused by population growth. The Japanese rice varieties maintained by Kobe University and used in this research could be used as valuable genetic resources to help identify other genes and breed new crop species.
Research team members for this study included Associate Professor YAMASAKI Masanori (Kobe University Graduate School of Agricultural Science Food Resources Education and Research Center), Professor MATSUOKA Makoto (Nagoya University Bioscience and Biotechnology Center), YANO Kenji (Nagoya University Bioscience and Biotechnology Center, currently research associate at the University of Tokyo Graduate School of Agricultural and Life Sciences), and YAMAMOTO Eiji (researcher at the NARO Institute of Vegetable and Tea Science).
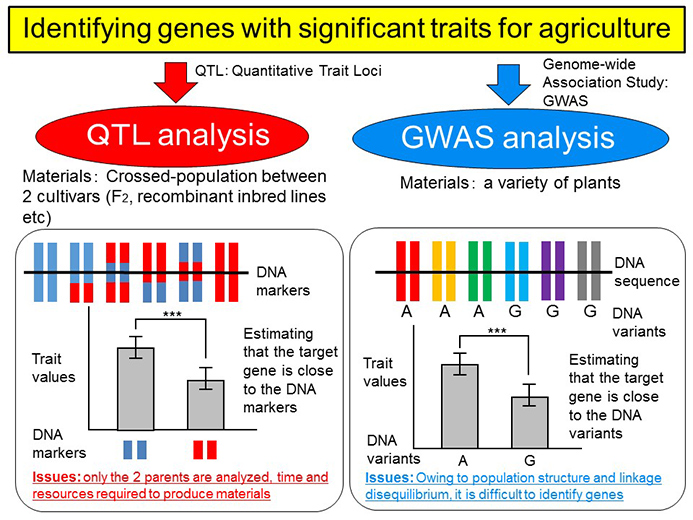
QTL analysis and genome-wide association study (GWAS). This time the team focused on GWAS.
Funding
Grant in Aid for Scientific Research from the Japan Society for the Promotion of Science; Council for Science, Technology and Innovation SIP (Cross-ministerial Strategic Innovation Promotion Program), and the Japan Society for the Promotion of Science
Journal information
- Title
- “Genome-wide association study using whole-genome sequencing rapidly identifies new genes influencing agronomic traits in rice.”
- Authors
- Kenji Yano, Eiji Yamamoto, Koichiro Aya, Hideyuki Takeuchi, Pei-ching Lo, Li Hu, Masanori Yamasaki, Shinya Yoshida, Hideki Kitano, Ko Hirano, and Makoto Matsuoka
- Journal
- Nature Genetics