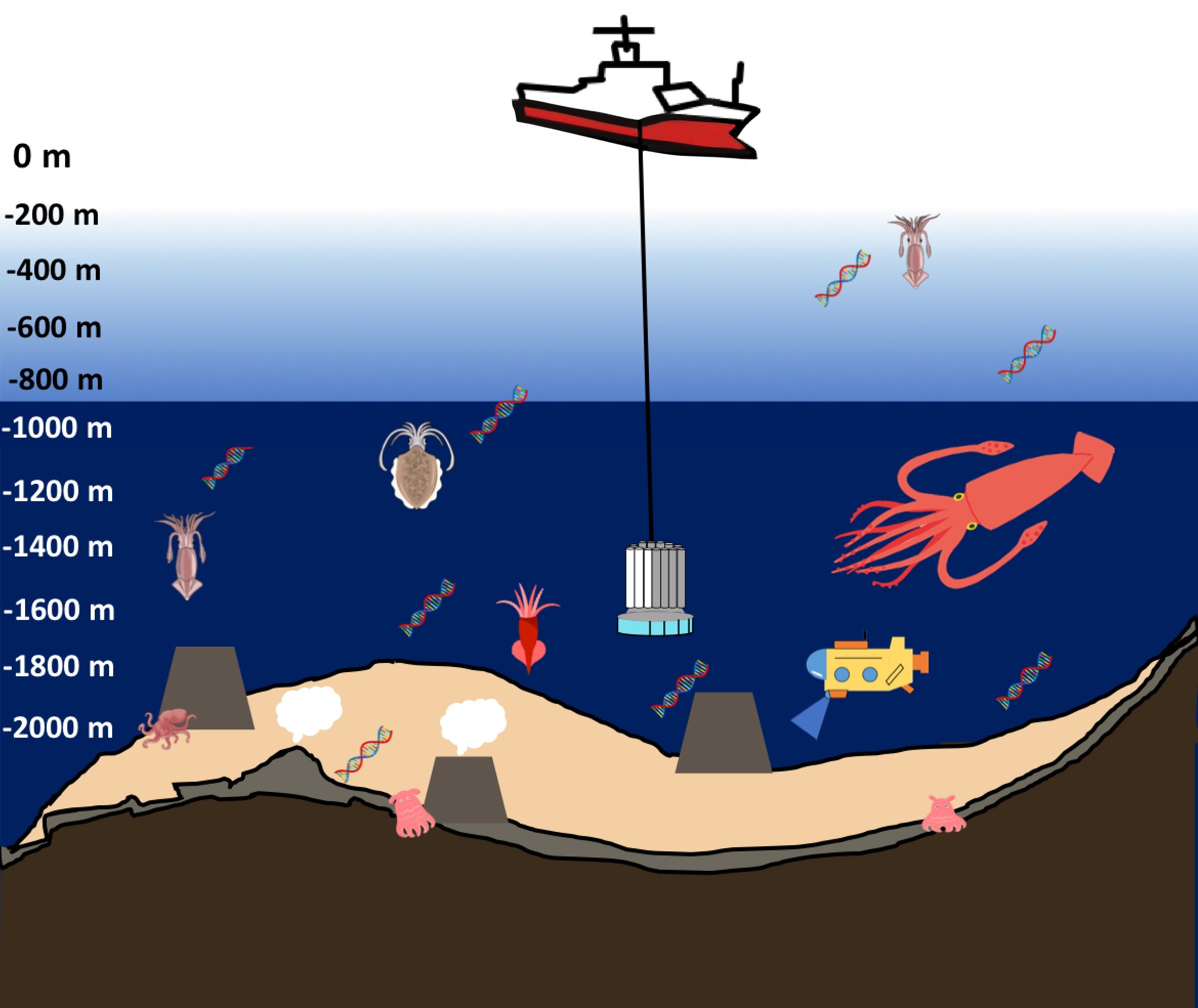
Environmental DNA surveys allow us to see what’s living in bodies of water like oceans, rivers and ponds by analyzing the DNA of just a cup of water taken from these sources. About a decade and a half has passed since the chance discovery of floating DNA of aquatic organisms came during other research, and since then, survey methods have been established and knowhow has been passed on to researchers from a wide range of universities and research institutes, as research continues to evolve and improve each day. We talked with Professor MINAMOTO Toshifumi of the Graduate School of Human Development and Environment, a pioneer in the field, about his journey in environmental DNA research, the current state of its social implementation and future prospects for the field.
What can a cup of water tell us now?
Specifically, what are environmental DNA surveys?
Minamoto:
Environmental DNA is the DNA of organisms found all over the environment, in things like water, soil and air. One easy example is fish: they release bodily waste, cells and even DNA itself into the water. The naked eye can’t see it, but a cup of water is full of the DNA of fish and other organisms.
We learned that there are detectable levels of this DNA in the environment a little over 15 years ago in 2008, when I was studying the koi herpesvirus, a virus that infects fish. As I was developing technology which collects water and counts the number of viruses in it, I happened to notice that this clean water had an enormous amount of carp DNA in it. I later learned that French researchers had published a paper about six months earlier reporting the detection of tadpole DNA in water. Of course, my discovery was the first in Japan.
A huge turning point for me was when our research was selected to the Japan Science and Technology Agency’s CREST Program. This was a large project in which we estimated the population of Japanese jack mackerel in Maizuru Bay using environmental DNA. During this research, I was also able to simultaneously collaborate in developing a method for listing which fish’s DNA was found in a cup of water. Now, this method is gaining some prominent attention.
Specifically, we mainly analyze DNA sequences of mitochondria found in cells. When mitochondrial DNA is inside a cell, it exists as a circular molecule of about 16,000 DNA base pairs in length, with a base being one of the four “letters” that make up a string of DNA, namely adenine (A), cytosine (C), guanine (G) and thymine (T). In most DNA analyses, researchers typically examine a region that is about 500-1,000 base pairs long.
On the other hand, environmental DNA found in the natural settings does not usually occur as an intact 16,000-base-pair molecule, but rather, it’s often fragmented into smaller pieces. In many cases, each fragment is even shorter than 500 base pairs, so it can’t be analyzed using conventional methods. As a result, we have to analyze pieces of DNA that are only around 100 or 150 base pairs long. In order to be able to analyze even DNA sequences that have been fragmented, we deliberately find a region for the base sequence with a low amount of information that we can identify to perform our analysis. From this environmental DNA, we can determine all the fish species present in the water.
Can you even determine how many individuals live there just by using survey data?
Minamoto:
If there are lots of fish, then the DNA released from them will also be relatively more abundant. The opposite is also true, meaning that the positive correlation between the amount of environmental DNA and number of fish in the water. Under exact same conditions, it would be possible to compare population sizes, but differences in season and location cause these conditions to change, which makes comparisons difficult. If we want to actually know the number of individuals that make up a population, environmental conditions would be different for each. Though, if you wanted to just broadly know whether there were “a lot” or a “a few” of a certain fish, I suppose you could still determine that.
It hasn’t yet been sufficiently determined what decides that. Fish are cold-blooded animals, which forces their metabolism to drop as water cools, in turn lowering the amount of DNA they discharge. The speed at which the DNA degrades also slows down in cooler water, so due to differences in water temperature, we don’t really know which environmental DNA is abundant and which is less so. Ultimately, the amount of environmental DNA is the difference between how much DNA was released and how much of it was degraded. There are also water quality factors outside of water temperature, like bacteria’s involvement in DNA degradation. The speed of DNA release and degradation change based on various parameters.
In our research counting mackerels in Maizuru Bay, we were really only able to accurately predict the number of individuals in a certain month and in a certain area of the ocean. One wouldn’t be able to simply apply the same method we used in other oceans. Different seasons would also lead to different results. Since there are many parameters influencing these results, it’s difficult to generalize this method, at least for the time being. For instance, if you wanted to see whether the number of fish have increased or decreased from year to year in the exact same sea during the exact same season, it should be possible to do that.

Full-scale adoption by the National Census on River and Dam Environments from next year
How much have you implemented these environmental DNA surveys into society?
Minamoto:
We’ve made significant progress in permeating environmental surveys. The Ministry of Land, Infrastructure, Transportation and Tourism conducts its National Census on River and Dam Environments over the course of five years, performing surveys on what animals live in class A rivers throughout Japan. In the 35 years since the survey Japan, they’ve made 7 full trips around the country, and from FY 2026, almost all surveys on fish will be conducted using environmental DNA surveys. Called “environmental DNA metabarcoding analysis,” this method has been recognized by the central government as being sufficiently useful for surveys.
Previously, we had to catch and record fish on site, but environmental DNA surveys have allowed us to save ourselves time and effort while greatly increasing the amount of data we receive. In terms of the number of survey locations, we’re able to cover about 10 times what we used to, which should greatly increase survey accuracy. This could become one of the largest environmental DNA surveys in the world; I’m so excited to see what kind of data we can acquire. We’re able to accurately survey both the presence and species of fish, which will allow us to investigate biodiversity in rivers. There are cases in which rare or invasive species can’t be found in capture-based surveys, but through environmental DNA surveys, we may find new habitats for rare or invasive species. It’s been over a decade since I learned of environmental DNA, and I think it’s incredible how much it has developed since then.
Environmental DNA surveys have even expanded overseas, with surveys actively carried out in European countries, the USA, Australia and New Zealand. Large countries should see large benefits from these surveys.
Assessing breeding status from environmental DNA
What kind of environmental DNA research are you undertaking now?
Minamoto:
One thing I’m working on is improving environmental DNA survey techniques. Environmental DNA isn’t foolproof, so there are instances in which a single cup of water isn’t enough to detect DNA from rare species. I’m conducting research that attempts to improve sensitivity by using methods that allow us to gather more data or use other DNA markers besides mitochondria.

In another research topic, we are now able to use environmental DNA to learn 1) what organisms are doing and 2) their condition. Specifically, we’ve developed a technique that shows us where or when animals are breeding. Up until this point, environmental DNA surveys have only told us about the existence of certain species. Taking the long-living Japanese giant salamander as an example, we weren’t able to assess whether or not there was a healthy generation turnover among the population. In conservation biology, it’s extremely important to ascertain breeding periods and locations, and we’re developing a technique that will allow us to do just that. We’ve already succeeded in capturing breeding signals from water collected in aquarium experiments in our laboratory and biotopes on campus.
As for specifically how we go about these surveys, DNA differs between animals in their normal state and animals in a breeding state in a process called “methylation.” We observe changes in cytosine (C), one of the bases in DNA sequences. Now we’re able to obtain information on methylation associated with environmental DNA bases that we were previously unable to gather simply by analyzing base sequences of environmental DNA surveys. We were the first in the world to establish this in vertebrates. This methylation information changes in response to an animal’s state. For instance, if we detect the DNA of the unique methylated state of sperm, we know that the sperm has been released, which reveals that breeding has taken place.
Initially, we’re focusing on breeding, but if we can improve techniques for analyzing methylation, I predict we can even learn animals’ age. Just as there are genetic regions where methylation states change during breeding, there are also regions that change according to age. I hope to find those areas and learn more about individual animals.
Once this aging structure is elucidated, we can then know whether the animals are undergoing a proper generation turnover. We may even be able to draw up a population pyramid that tracks whether there are larger numbers of younger or older individuals within the population. If we can figure out which populations in Japan are aging and where these populations are going through a healthy generation turnover, we can then ascertain where we need to prioritize our conservation efforts. I’d like to spend the next 10 years or so performing research on this topic.
Tell me about how these methods have been adopted by research institutes. Are there any applications that you didn’t expect?
Minamoto:
Environmental DNA research originally began with me and my colleagues, but now it’s spread to research institutes and universities all over Japan. In April 2018, we established the eDNA Society (288 regular members and 36 supporting member organizations as of the end of August 2025) and are currently carrying out research there as well. There haven’t been many unexpected use cases, but one that I found interesting was research on infectious diseases.
More than half of infectious diseases are zoonotic, with shared pathogens between wild animals and humans. There are, however, many infectious diseases in which we don’t know the pathogen’s “natural host,” so it’s unclear which animals possess these pathogens.
Previously, I performed joint research with the University of the Ryukyus in order to identify the “culprit.” Each year, there is a survey on a pathogen called Leptospira bacteria which causes diseases in the rivers in Okinawa. We didn’t know which animals living around rivers release this Leptospira bacteria into the water. After collecting water many times and analyzing environmental DNA, we found that a certain animal’s DNA appeared whenever the DNA of the pathogen appeared in our analyses, allowing us to assume who the “culprit” was. This way of thinking will reveal the relationship between organisms. Ultimately, infectious disease is a mutual interaction between wildlife and pathogens, so if we can understand the relationship between these organisms, it may allow us to stop the cycle of disease in the future.
Are human actions “nature positive”?
Tell me about your plans for future research and other activities.
Minamoto:
Just as the National Census on River and Dam Environments has shown, environmental DNA, which provides a list of animals living in a given area, has transcended the realm of research and is now being implemented into society. What I’d like to pursue now is research on how these animals live.

Also, I think research will move towards advancing what we are able to understand when we use techniques implemented in society that collect large amounts of data. Recently, a way of thinking that’s gathering attention is “nature positive.” The aim is to halt and reverse the loss of biodiversity, but in actuality, it’s difficult to measure whether what’s being done is really nature positive. For instance, if you continue to keep a database on the National Census on River and Dam Environments for decades, you would be able to know whether biodiversity is increasing or decreasing. Or if you built a structure like a dam, you would be able to know what happened to biodiversity in the area. By continuing to conduct environmental DNA surveys, we should be able to elucidate the positive or negative effects human activity has on nature.
You could say the same about global warming. We’ve now been carrying out environmental DNA sampling of the oceans from Kobe to Awaji Island and even down to Shikoku for the past 10 years or so, but if we were to continue, we might be able to see what kinds of effects recent global warming and the large meander of the Kuroshio Current have had on the waters around Japan. Gathering data over a long period of time should lead to an even greater understanding of organisms than the one we have now. This means that we would be handling some really, really big data, so we would need to integrate some data science as well.
Countries from around the world have collected a large amount of environmental DNA from all kinds of locations, so it’s important that we have an international standard for this data in order to make proper comparisons. Currently, researchers have begun to undertake this endeavor at the volunteer level, and the eDNA Society, where I serve as the president, is also fully committed to these efforts. I think that data collected using standardized methods by researchers from around the world should advance our understanding of how ocean warming has affected the distribution of the global fish population. We weren’t ever able to do this before environmental DNA.
The potential of environmental DNA surveys that transcend species
You’ve been carrying out research on developing environmental DNA analysis techniques that apply to all species and establishing indexes for nature positivity in the "Habataku" Project for Next-Generation Interdisciplinary Co-creation. What does that involve?
Minamoto:
When conducting environmental DNA surveys, we need to perform analysis experiments that differ depending on the animal, i.e., fish analyses for fish and amphibian analyses for amphibians, and we have to perform these different experiments many, many times. I want to develop a method in which we can determine what species live in a location by performing a single analysis by thoroughly analyzing the DNA in a single sample. We currently don’t have any established indexes for nature positivity, but I think that by combining it with this new environmental DNA technique, we can create just that. I’m trying to create these nature positivity indexes by collaborating with environmental economics researchers in the project and combining all kinds of data, like changes in fish, amphibian and insect populations.
Is there anything you want to say about activity outside of your research?
Minamoto:
At the Graduate School of Human Development and Environment, we have an organization called the “Kobe University Science Shop.” This organization connects with groups like the general public and high school students to deliver knowledge and technology held by the university. The organization also holds science cafes and public lectures and provides support for high school students who want to perform research. Moving forward, I’d like to gather more participants and expand our activity.
Resume
In March 1997, graduated from the Faculty of Science, Kyoto University. In March 1999, completed the master’s program, and in March 2003, received his doctoral degree in science at the Graduate School of Science, Kyoto University. In April 2003, became research associate at the Center for Ecological Research, Kyoto University. In April 2005, became post-doctoral researcher at the National Institute of Advanced Industrial Science and Technology. In April 2007, became senior project researcher at the Research Institute for Humanity and Nature. In November 2012, became project assistant professor, in October 2017, became associate professor and in July 2021, became professor at the Graduate School of Human Development and Environment, Kobe University