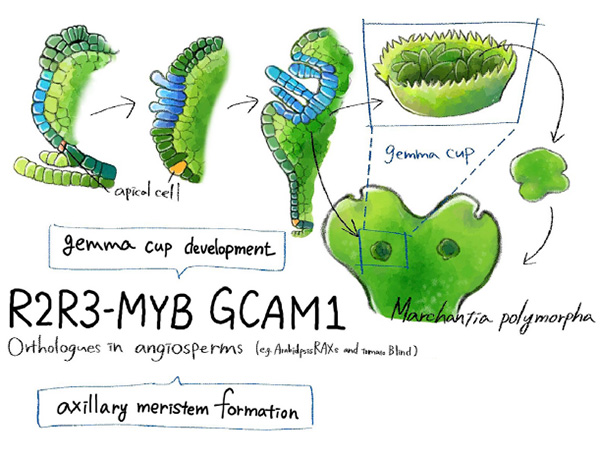
The mechanism by which liverwort (Marchantia polymorpha) asexually reproduces via the development of clonal progenies (gemmae) has been revealed by a cross-institutional research group. They discovered the gene ‘KARAPPO’, which is essential for initiating gemma development in liverwort. These findings are expected to contribute fundamental knowledge towards technological developments to boost agricultural and horticultural cultivation efficiency.
The collaborative research team consisted of members from institutions including Kobe University, Kyoto University, the National University of Singapore and the National Institute for Basic Biology, among others. Members from Kobe University’s Graduate School of Science included Professor Kimitsuke Ishizaki, Researcher Yukiko Yasui and Takuma Hiwatashi (a third year PhD student).
The results of this study were published in the American Journal ‘Current Biology’ on October 10, 2019.
Background
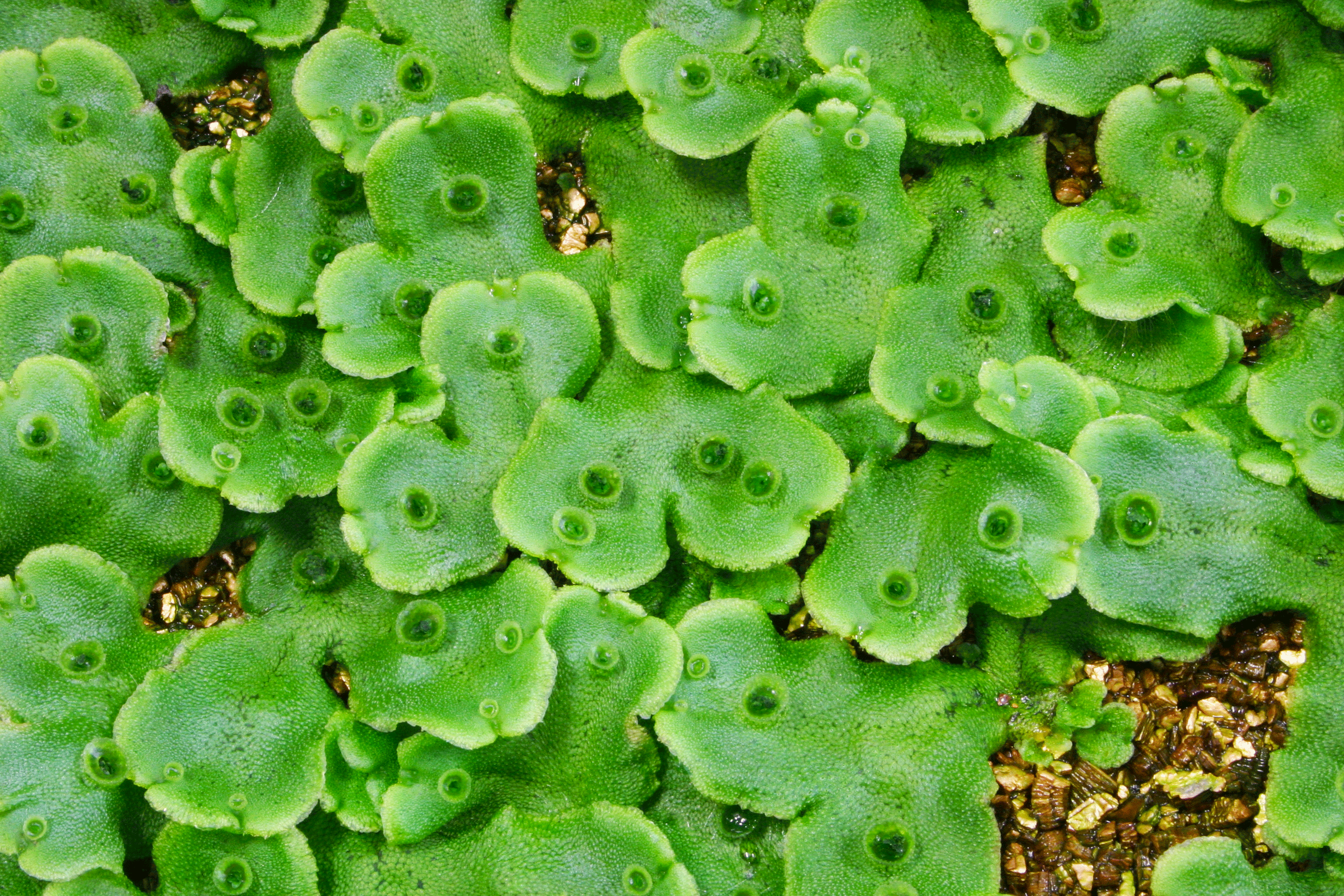
Vegetative reproduction is a form of asexual reproduction in which individual plants are developed directly from the tissues of the parent plant. Liverwort reproduces by forming clones of itself (collectively called gemmae) in a gemma cup (Figure 1). These cups form on the thallus, or body, of the plant. Inside the gemma cup, epidermal cells (that form a protective layer over the outside of a plant) undergo cell elongation followed by two cycles of asymmetrical cell division in order to form a gemma cell and a basal cell. This gemma cell continues to divide before finally forming a new plant.
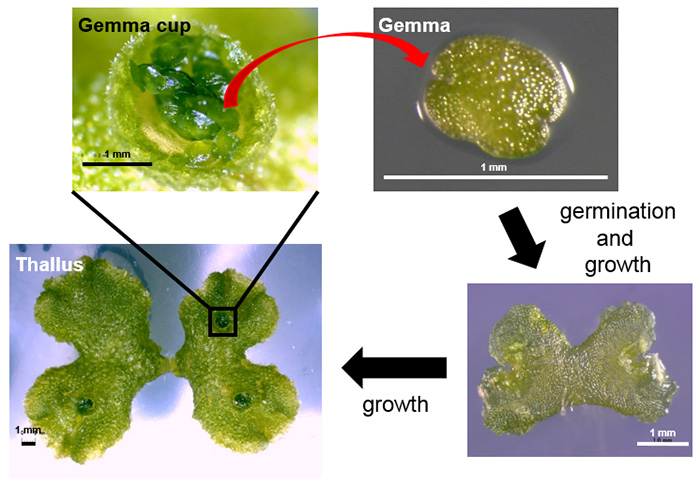

It is thought that bryophytes (liverworts, mosses, and hornworts) evolved from land plants’ algal ancestors more than 430 million years ago. Liverwort was one of the earliest diverging plants to develop characteristics allowing it to live on land. The whole-genome sequence information for liverwort has been available since 2017. The plants only carry a minimum set of genetic information- which make them useful for studying the effects of genetic modification and for helping scientists understand more about the evolution of mechanisms in land plants.
Methodology
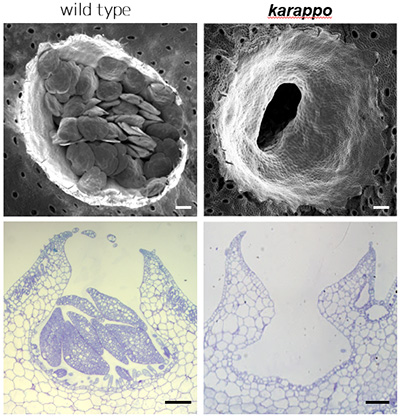
Above: Gemma cups as shown through a SEM (Scanning Electron Microscope)
Below: Cross section of the gemma cups
Scale bar: 100µm
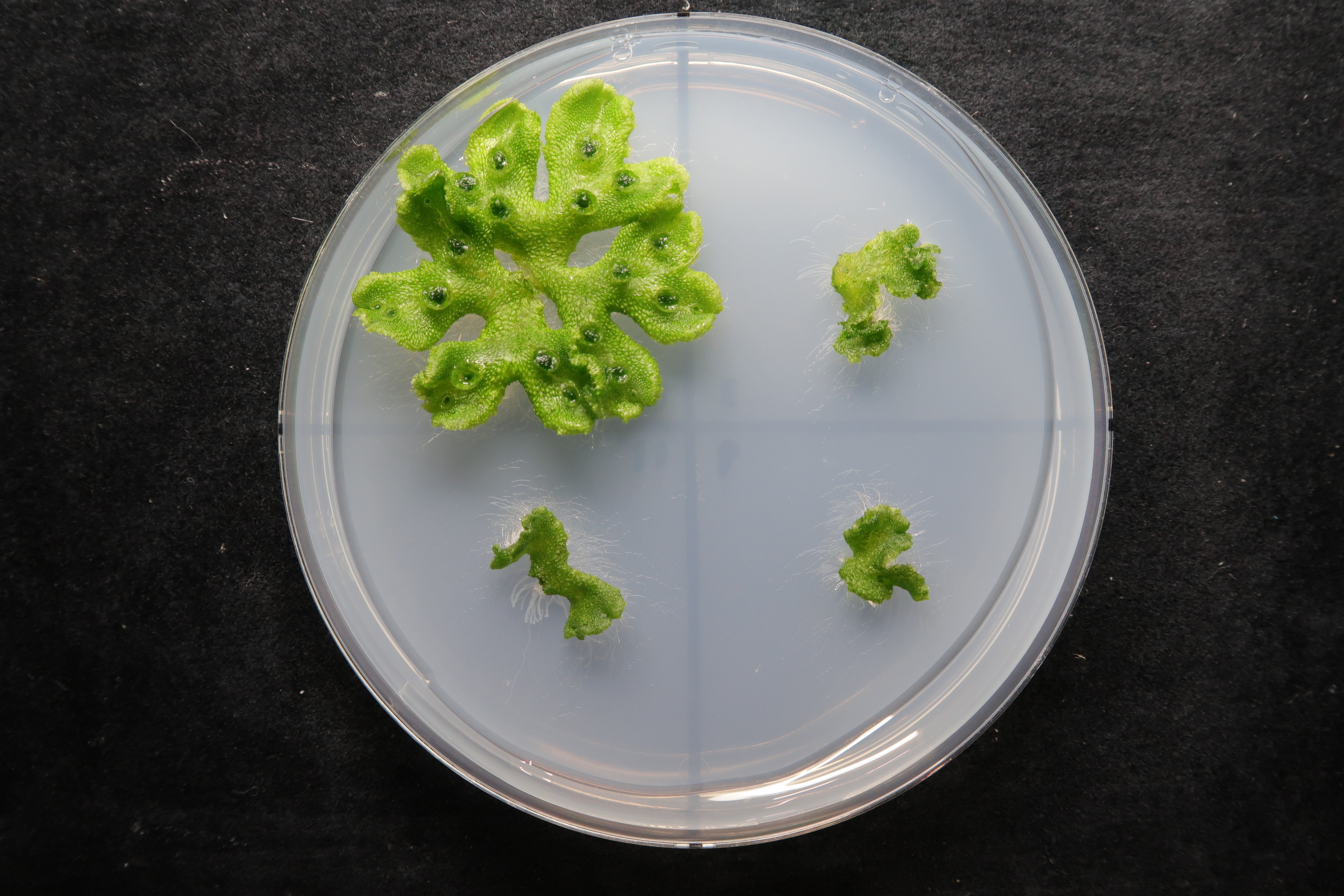
The research team focused on two mutant liverworts named karappo-1 and karappo-2. Karappo means empty in Japanese, and the mutants were given these names because no initial gemma developed in the cups on these plants (Figure 2). Next generation DNA sequencing of the mutants allowed the researchers to identify the gene responsible for gemmae development- which they named ‘KARAPPO’.
Subsequent analysis of the amino acid sequences revealed that the KARAPPO gene encodes the RopGEF. ROP is a type of small GTP-binding protein found in plants. It acts like a ‘switch’ and signals a range of cellular processes. ROP activation depends on guanine nucleotide exchange factors (GEFs) which catalyze the release of GDP to facilitate the subsequent binding of GTP in ROP. The RopGEF encoded by KARAPPO was found to trigger cell elongation and asymmetrical cell divisions in the initial stage of gemma development. This demonstrated that the KARAPPO gene is absolutely essential in triggering the processes for vegetative reproduction in liverwort (Figure 3).
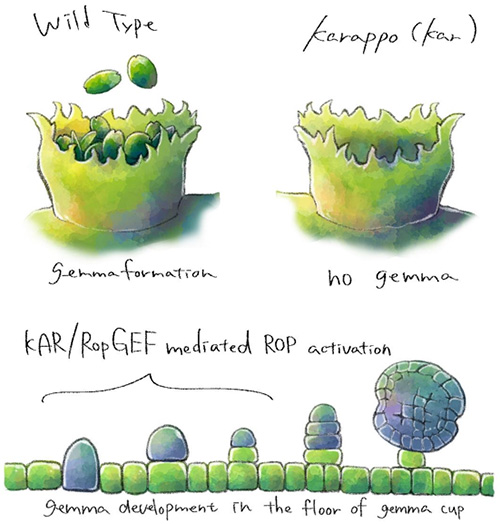
Conclusion:
These research results have illuminated fundamental knowledge regarding vegetative reproduction mechanisms. The findings highlight the importance of ROP-driven asymmetric division of differentiated cells in land plants. The development of this mechanism allowed land plants to asexually produce and regenerate clones, allowing them to dominate the terrestrial ecosystem. Further research into these mechanisms could result in revolutionary technological developments in agriculture and biotechnology.
Journal information
- Title
- “The RopGEF KARAPPO is Essential for the Initiation of Vegetative Reproduction in Marchantia polymorpha”
- DOI
- 10.1016/j.cub.2019.08.071
- Authors
- Takuma Hiwatashi1, Honzhen Goh2, Yukiko Yasui1,3, Li Quan Koh2, Hideyuki Takami1, Masataka Kajikawa3,4, Hiroyuki Kirita3, Takehiko Kanazawa5,6, Naoki Minamino5, Taisuke Togawa4, Mayuko Sato7, Mayumi Wakazaki7, Katsushi Yamaguchi8, Shuji Shigenobu8, Hidehiro Fukaki1, Tetsuro Mimura1, Kiminori Toyooka7, Shinichiro Sawa9, Katsuyuki T. Yamato4, Takashi Ueda5,6, Daisuke Urano2, Takayuki Kohchi3, and Kimitsune Ishizaki1*
1 Graduate School of Science, Kobe University
2 Temasek Life Sciences Laboratory, National University of Singapore
3 Graduate School of Biostudies, Kyoto University
4 Faculty of Biology-Oriented Science and Technology, Kindai University
5 Division of Cellular Dynamics, National Institute for Basic Biology (NIBB)
6 Department of Basic Biology, The Graduate University for Advanced Studies (SOKENDAI)
7 RIKEN Center for Sustainable Resource Science.
8 Functional Genomics Facility, National Institute for Basic Biology (NIBB)
9 Graduate School of Science and Technology, Kumamoto University
*Corresponding author - Journal
- Current Biology