A collaborative research group has succeeded in identifying an important transcription factor, GCAM1, which allows liverwort plants to asexually reproduce through creating clonal progenies (vegetative reproduction). Furthermore, this transcription factor was revealed to have the same origin as those which regulate secondary bud formation in angiosperms. That this transcription factor was found in liverwort suggests that it was an important ancient development in the common ancestor of land plants at least 430 million years ago.
It is hoped that these discoveries will lead to the development of technologies to increase the cultivation rate of a variety of plants in agriculture and horticulture.
The group consisted of researcher Yukiko Yasui (now an assistant professor at Kyoto University) and Associate Professor Kimitsune Ishizaki from Kobe University’s Graduate School of Science, in addition to members from Kyoto, Shinshu and Kindai Universities, and the Max Planck Institute for Plant Breeding Research (MPIZ).
These research results were published in the American scientific journal ‘Current Biology’ on November 7, 2019.
Introduction
Meristems are pools of undifferentiated, or unspecialized cells located in areas of plants where growth can occur- usually at the tips of the plant. These meristem allow for new shoots and flowers to grow, in addition to enabling many varieties of plants to reproduce asexually by creating clones of themselves. This process is called vegetative reproduction. The clonal progenies formed via vegetative reproduction have the same genetic makeup as the parent plant. Vegetative reproduction occurs in different areas depending on the variety of plant, for example, stems (potato, strawberry), roots (sweet potato) or even leaves (kalanchoe). It can also be induced using a number of techniques, such as cuttings and grafting, and is commonly utilized to produce plants in both agriculture and horticulture. However, many aspects of the mechanism behind this reproduction system remain unknown.
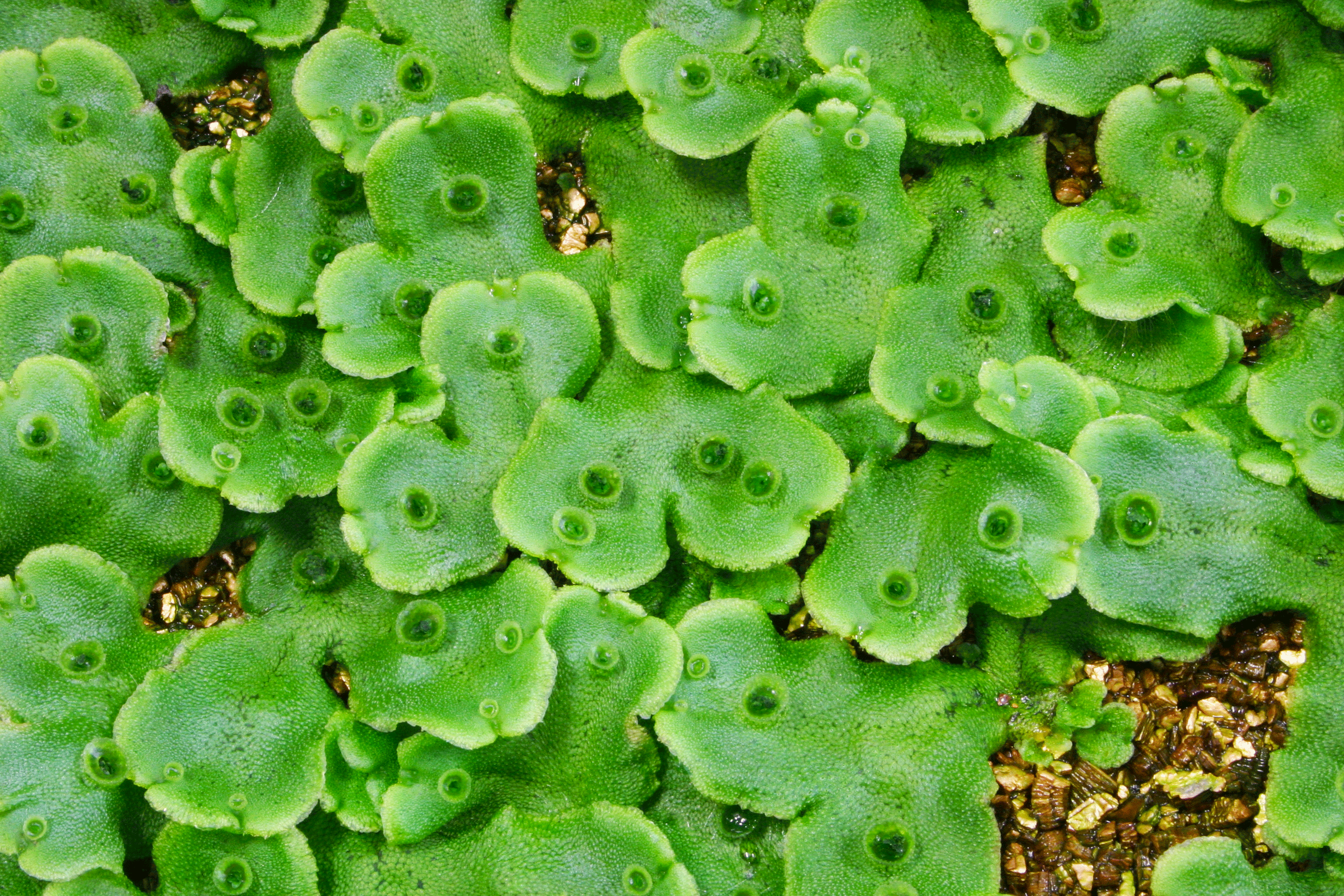
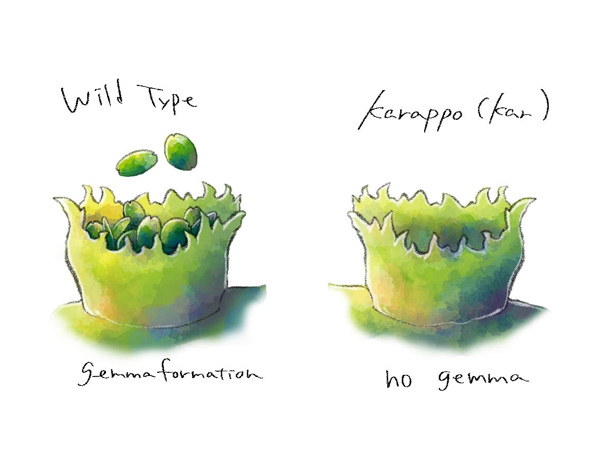
Vegetative reproduction in liverwort takes place inside gemma cups, which form on the surface of the body of the plant (Figure 1). A research group consisting of some of the same members recently illuminated the mechanism by which gemmae develop within the gemma cup. However, it was still unclear what mechanism allowed gemma cups to form in the first place.
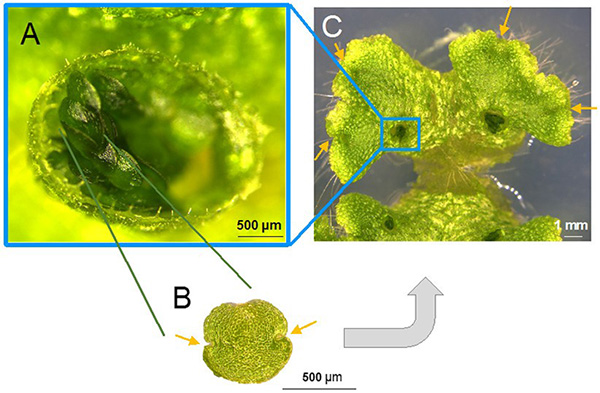

A. A liverwort gemma cup. The gemma (clone) inside is visible.
B. Enlarged image of the gemma. There are meristem on both sides (as indicated by the arrows), which contain stem cells.
C. Photo of gemma growth in cups on the surface of the plant. Divergence and growth in the meristem area is indicated by the arrows.
Research methodology
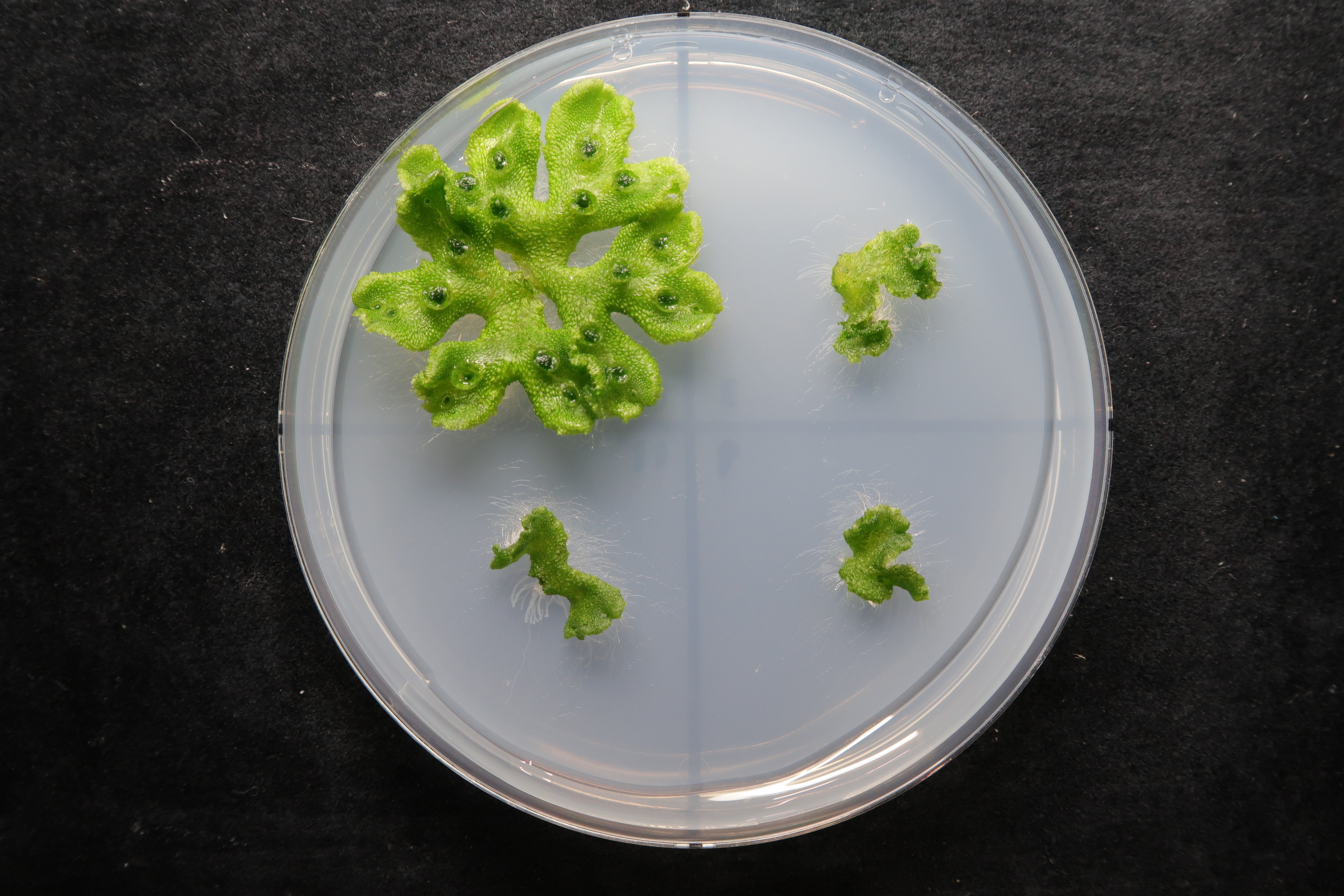
This research group sought to illuminate which genes play a large role in gemma cup formation via comprehensive analysis of gene expression. Through this analysis they found one transcription factor, which they called Gemma cup-associated MYB1 (GCAM1). They discovered that removing this GCAM1 gene from the plant prevented gemma cups from forming (GCAM1 mutant- Figure 2). However the removal of this gene didn’t affect other aspects of the plant’s growth apart from the gemma cups, leading them to conclude that the GCAM1 proteins that encode the GCAM1 gene are vital for triggering the formation of gemma cups.
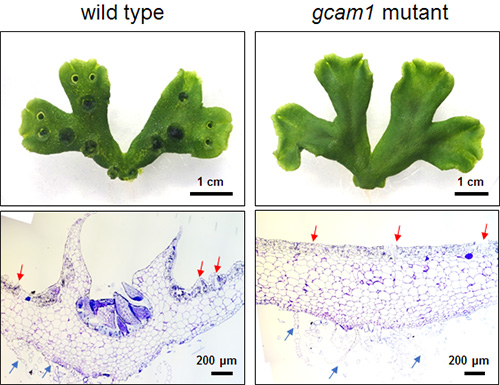
Above: Three week old liverwort plants. On the left is a regular wild type liverwort (used as the control) and on the right is a mutant liverwort with the GCAM1 gene removed.
Below: Cross-section of the gemma cup. The red arrows indicate air chambers and the blue arrows show the abdominal scale. In the mutant (on the right), the gemma cups didn’t form.
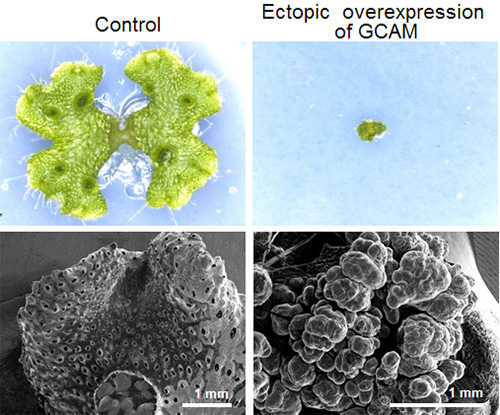
Above: 2 week old liverwort plants
Below: SEM (scanning electron microscope) image
Next, they illuminated the function of the GCAM1 proteins with the aim of controlling gemma cup formation. To do this, they created an overexpression of GCAM1 proteins inside the liverwort plant. When GCAM1’s protein function is overexpressed, the plant is unable to grow and there is an increase in undifferentiated cells with stem cell properties (Figure 3). Based on this result, it is thought that GCAM1 is responsible for suppressing cell differentiation in the meristem regions, which under normal conditions would allow for gemma cup and gemmae formation.
GCAM1 is an R2R3-MYB transcription factor gene. The same subfamily of genes in Angiosperms is known to have an axillary bud forming function (RAXs in Arabidopsis thaliana and Blind in tomato plants). To investigate this evolutionary relationship, an experiment was carried out using the model plant Arabidopsis thaliana.
It has been previously discovered that mutant Arabidopsis thaliana plants without the RAXs gene develop few axillary buds compared to the wild-type (control) plant. However, this research group found that introducing the GCAM1 gene into this mutant caused the number of axillary buds to increase. Liverwort’s GCAM1 gene displayed the same function in Arabidopsis thaliana, indicating that the RAXs gene and the GCAM1 gene are orthologs (Figure 4). Orthologs are genes in different species that evolved from a common ancestral gene and thus retain the same function.
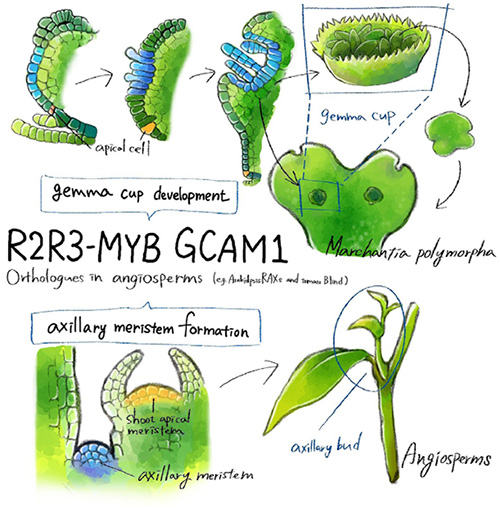
The GCAM1 transcription factor gene, which forms gemma cups in liverwort is an ortholog of that which controls axillary bud formation in angiosperms.
Conclusion
It was revealed that liverwort’s GCAM1 gene and the genes that are responsible for axillary bud formation in angiosperms are orthologs. From this finding, it is supposed that the gemma cup formation in liverwort and axillary bud formation in angiosperms have the same mechanism.
Technology to control axillary shoot formation in angiosperms could be used to directly increase crop production. It is hoped that further research into GCAM1 could provide a foundation for such technology. Next, it is necessary to illuminate the network involved in GCAM1 management, and to investigate the commonalities between liverwort and angiosperms. This will contribute not only to the understanding of vegetative reproduction in liverwort but will also reveal the common mechanism by which plants produce new buds.
Glossary:
- Liverwort (Marchantia polymorpha)
- propagates via vegetative reproduction- whereby clones (gemmae) of the parent plant develop inside gemma cups which form on the body of the plant. The whole genome information for liverwort has been available since 2017 and the plant has received attention as a model for research into the evolution of mechanisms in land plants and gene functions.
- Meristem
- Tissue containing undifferentiated cells. It is found in parts of the plant where growth can occur.
- Arabidopsis thaliana
- Also known as thale cress, it is a small flowering plant (angiosperm) in the brassicaceae (mustard) family. The whole genome sequence for this plant has been available since 2000, meaning that it is often used as a model in plant biology research studies investigating gene functions and mechanisms.
- Angiosperms
- are flowering plants that have seeds enclosed within a carpel. Many kinds of land plant are classified as angiosperm- including herbaceous plants, shrubs and most trees.
- Transcription factors
- are proteins that help turn specific genes (and their functions) ‘on’ or ‘off’.
- Undifferentiated cells
- Basic cells that have yet to develop any specialized functions.
Journal information
- Title
- “GEMMA CUP-ASSOCIATED MYB1, an Ortholog of Axillary Meristem Regulators, Is Essential in Vegetative Reproduction in Marchantia polymorpha”
- DOI
- 10.1016/j.cub.2019.10.004
- Authors
- Yukiko Yasui,1,2# Shigeyuki Tsukamoto,1,# Tomomi Sugaya,3 Ryuichi Nishihama,2 Quan Wang,4 Hirotaka Kato,1 Katsuyuki T. Yamato,5 Hidehiro Fukaki,1 Tetsuro Mimura,1 Hiroyoshi Kubo,3 Klaus Theres,4 Takayuki Kohchi,2 and Kimitsune Ishizaki1*
1 Graduate School of Science, Kobe University
2 Graduate School of Biostudies, Kyoto University
3 Faculty of Science, Shinshu University
4 Max Planck Institute for Plant Breeding Research (MPIZ)
5 Faculty of Biology-Oriented Science and Technology, Kindai University
*Corresponding author - Journal
- Current Biology