The roles of histone modification (*1), including H3K4me3 and H3K36me3, in the regulation of gene expression in Chinese cabbage plants have been elucidated.
This discovery was made by an international research group, which included:
- Hasan Mehraj (a PhD. student at Kobe University’s Graduate School of Agricultural Science)
- TAKAHASHI Satoshi (technical staff at RIKEN Center for Sustainable Resource Science)
- Associate Professor FUJIMOTO Ryo (Kobe University’s Graduate School of Agricultural Science)
- SEKI Motoaki (team leader at RIKEN Center for Sustainable Resource Science)
- Elizabeth S. Dennis (of the Commonwealth Scientific and Industrial Research Organisation (CSIRO), Australia)
These research results were published online in Frontiers in Plant Science on June 7, 2021.
Main points
- Histone modification is important for the regulation of gene expression in Chinese cabbage. The researchers revealed the roles that the tri-methylated histone H3 lysine 4 (H3K4me3) and H3K36me3 play in this process.
- H3K36me3 is important for activating gene expression and ensuring consistent gene expression between different tissues.
- The researchers identified genes with bivalent chromatin structures that have both transcription activating (H3K4me3) and repressing histone modifications (H3K27me3).
Research Background
Differences in gene expression levels have been observed between individuals whose DNA sequence consists of the same genes and even within the body of a single individual. For example, even within the same plant there are different structures in which different groups of genes are expressed, such as leaves, roots and flowers. In this case, the nucleotide sequence of the DNA is the same in each structure, therefore information other than the DNA sequence regulates gene expression. This kind of regulation of gene expression that is not determined by the DNA sequence is called epigenetic (*2) regulation. Known examples of this include DNA methylation and histone modification. With the recent advances in technology to determine DNA sequences, it has become possible to determine the DNA sequence for the entire genome of various species. Consequently, genome-wide epigenetic modification patterns (epigenome *3) have been mapped for various species.
The whole genome for Chinese cabbage (Brassica rapa L.), which is the subject of this research study, was sequenced in 2011. This research group has previously published findings on the distribution of H3K27me3 modification (a genome-wide repressive histone modification), however they had not reported on other histone marks such as H3K4me3 and H3K36me3 (Figure 1). In addition, they had not addressed the relationship between the transcription activating histone marks H3K4me3 and H3K36me3, and the transcription-repressing H3K27me3.
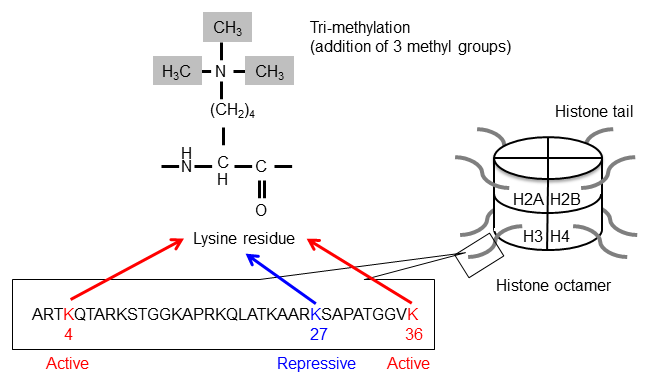
The tri-methylation of the 4th lysine residue (K) of Histone H3 results in the H3K4me3 modification. H3K36me3 and H3K27me3 result from the tri-methylation of lysine 36 and lysine 27, respectively. The sequence of a histone tail and the modification of the lysine residues is shown.
Research Methodology
Using leaf tissue, the researchers of this group defined the state of H3K4me3 and H3K36me3 modifications at the whole genome level in two lines of Chinese cabbage. In the two lines, they compared the average value of expression levels for genes with H3K4me3 marks (around 17,000 genes) and H3K36me3 marks (around 12,000 genes) with the average expression levels for all of the genes (approximately 40,000). The results revealed that expression levels were comparatively higher for the genes with these histone modifications, and this trend was particularly observable in genes with H3K36me3 (Figure 2). This indicates that H3K4me3 and H3K36me3 modifications are responsible for activating gene expression. Furthermore, it was also revealed that expression levels for H3K4me3 and H3K36me3-marked genes have a tendency to be resistant to change, which is indicative of constitutive gene expression (Figure 3).
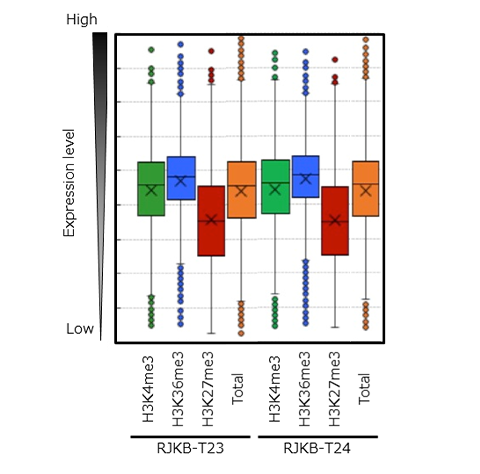
‘Total’ indicates the expression levels for all genes.
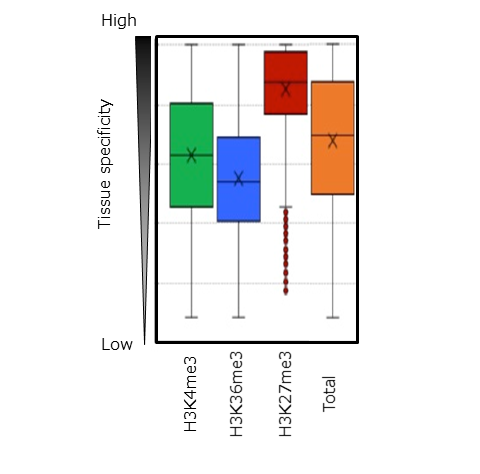
‘Total’ indicates the expression levels for all genes.
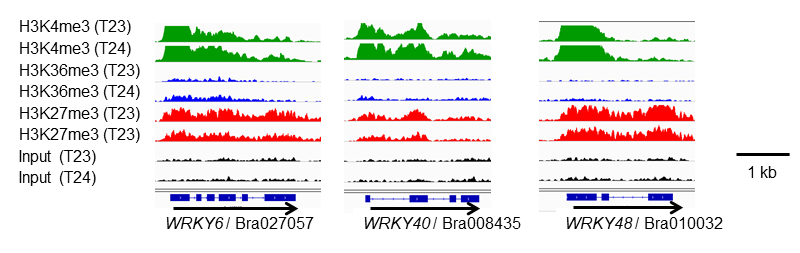
Building upon their previously published research on transcription repressing H3K27me3, the researchers investigated the relationships between the 3 histone modifications and found that H3K36me3 and H3K27me3 have a competitive relationship with each other. In addition, they also discovered a group of around 3,700 genes that could have bivalent chromatin structures with both H3K4me3 and H3K27me3 modifications (Figure 4).
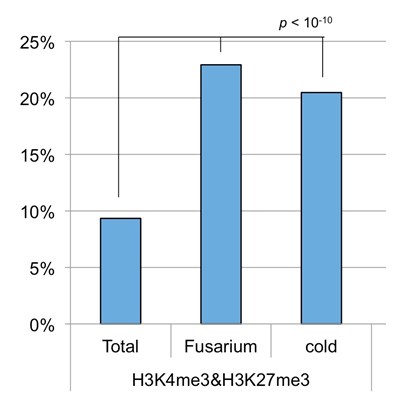
Biotic stress: Genes that were expressed differently after being infected with the pathogenic fungus Fusarium. Environmental stress: Genes that were expressed differentially after 4 weeks exposure to cold. Total: The percentage of genes with H3K4me3 and H3K27me3 marks out of the total number of genes.
Among the group of genes that have a bivalent chromatin structure, many were found to encode transcription factors. Furthermore, numerous genes in the group changed their expression in response to biotic and environmental stress (Figure 5). This suggests the possibility that bivalent chromatin structures may form in circumstances where both activation and repression of gene expression is needed. In fact, the researchers used a method to detect bivalent chromatin structures and confirmed that this type of structure definitely forms in a number of genes.
This research has revealed the role of active histone modifications (H3K4me3 and H3K36me3) in gene expression, in addition to their relationship with repressive histone modifications (H3K27me3).
Further Research
The results of this research have illuminated the genome-wide role of H3K4me3 and H3K36me3 modifications in Chinese cabbage plants. Next, the group will conduct a detailed investigation into the relationship between these histone modifications and the agricultural traits of Chinese cabbage by clarifying the roles of H3K4me3 and H3K36me3 in individual genes and their connection to plant phenotype. In the current study the researchers also identified genes with a bivalent chromatin structure that contain both active and repressive modifications, suggesting that these genes play an important role in the plants’ response to stress. Subsequently, they also plan to investigate these genes on an individual level to understand exactly how they are related to Chinese cabbage plants’ stress response.
Glossary
- *1 Histone modification
- Histones are proteins found inside the nucleus. They are arranged in a histone octamer structure, which is an eight protein complex consisting of two copies of each of the four core histone proteins (H2A, H2B, H3 and H4). The smallest unit of a chromosome is called a nucleosome and it consists of a section of DNA wrapped around a histone octamer. Multiple nucleosomes make up a chromatin structure. A histone modification is when a particular histone’s lysine base is acetylated or methylated, which changes the chromatin structure and causes gene expression to be activated or repressed.
- *2 Epigenetic
- Stable, heritable gene expression or transformation which originates from chromosomal alterations and doesn’t involve changes to the DNA sequence.
- *3 Epigenome
- A set of chemical compounds that attach to the DNA to modify its function without changing the sequence of the DNA. These marks change the way cells use the DNA’s instructions and include DNA methylation, histone modification, non-coding RNA and higher-order structures of chromatin.
Acknowledgements
This research received support from the following organizations:
- The Ministry of Education, Culture, Sports, Science and Technology (MEXT), KAKENHI grant (no. 221S0002).
- The Japan Society for the Promotion of Science (JSPS)’s Bilateral Joint Research Projects (Open Partnership Joint Research Projects).
- Fund for the Promotion of Joint International Research (JSPS, Grant number: 16KK0171).
- Also from JSPS, A Grant-in-Aid Young Scientists (A) (Grant number: 15H05614), and Grants in Aid for Scientific Research (B) (Grant numbers 18H02173 and 19H02947).
Journal Information
- Title
- “Characterization of histone H3 lysine 4 and 36 tri-methylation in Brassica rapa L.”
- DOI
- 10.3389/fpls.2021.659634
- Authors
- Hasan Mehraj+, Satoshi Takahashi+, Naomi Miyaji, Ayasha Akter, Yutaka Suzuki, Motoaki Seki, Elizabeth S. Dennis, Ryo Fujimoto
+ These authors equally contributed to this work - Journal
- Frontiers in Plant Science











