Many virus-derived DNA sequences can be found in the eukaryotic genomes of mammals, including those of humans. It has long been thought that these viral sequences defend the host against the corresponding viruses, but the exact mechanism by which they do this is unknown. Recent studies have shown that the mechanism is mediated by the latest class of RNA interference: piRNA.
piRNAs derived from integrated viral sequences may enact transgenerational antiviral immunity in eukaryotes, possibly in the same manner as CRISPR RNA in prokaryotes. The mechanism may represent a novel antiviral innate immunity in eukaryotes. In other words, this mechanism may allow offspring to inherit virus immunity from their parents.
These important findings have been reviewed and postulated by a collaborative group consisting of Dr. Youdiil Ophinni and Prof. Yoshitake Hayashi from Kobe University Graduate School of Medicine, team leader Nicholas F. Parrish from RIKEN, and Umberto Palatini from University of Pavia.
The paper was published in the November issue of the top-ranked review journal, Trends in Immunology, under the Cell Press publishing group.
Introduction
A portion of the eukaryotic genome, including about 8% of human DNA, is made from virus-derived sequences called endogenous viral elements (EVEs). Thought to be junk DNA at first, it has now been recognized that EVEs play an important role in many mammalian functions, although little was understood about their role in immunity. However, recent studies have indicated that EVEs may engender RNA-mediated immunity against corresponding viruses through the action of piRNA.
piRNA was first discovered in 2006 in the fruit fly Drosophila. It is a class of 24-31 nucleotides (nt) RNAs that bind with the PIWI protein and guide them to silence target RNA or DNA. It is recognized that piRNA-PIWI complexes target and silence transposable elements (TEs), the parasitic jumping gene found in abundance within the genome.
piRNAs are transcribed from specific loci called piRNA clusters that are widespread within the eukaryotic genome. The long piRNA precursors are processed by a family of PIWI proteins, producing mature piRNAs bound to PIWI. piRNA-PIWI complexes can then cleave target RNA in the cytoplasm, or enact histone or DNA methylation to target DNA in the nucleus.
Methodology
In this review paper, the authors compiled the results of studies within the last five years to postulate a hypothesis: that EVE-derived piRNA-PIWI may enable heritable, sequence-specific antiviral immunity in eukaryotes.
One important piece of evidence supporting this hypothesis is the enrichment of EVEs within piRNA clusters in many arthropods and mammals. EVEs consisting of endogenous bornavirus-like nucleoprotein elements (EBLNs) are enriched in piRNA clusters in many mammals including humans. They generate piRNAs that are antisense, or complementary, to the ancient bornavirus nucleoprotein mRNA. EVEs are also enriched within the piRNA clusters of many arthropods, and produce piRNA antisense to the corresponding arboviral RNA.
The strongest evidence is shown in domestic chickens, in which the horizontal spread of avian leukosis virus (ALV) is inhibited by ALV-derived EVE. ALV EVE also produces bona fide ALV-targeting piRNAs. A similar observation was made in koalas; EVE loci of koala retroviruses produce piRNA that may target and potentially silence the exogenous virus.
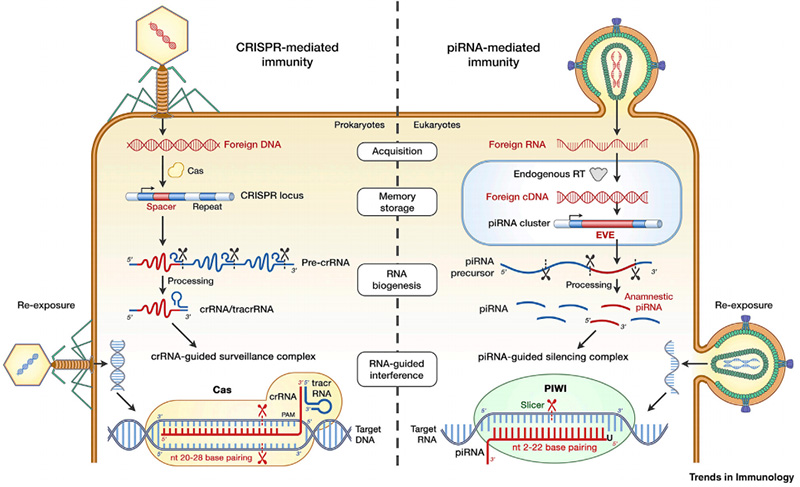
The authors postulated that horizontally spread viral sequences are integrated into the host genome to create EVE, which is inherited by progeny. EVEs located inside piRNA clusters produce piRNAs that may target and silence any corresponding viral infections. The authors further draw comparisons between this piRNA-PIWI antiviral immunity in eukaryotes to CRISPR-Cas in prokaryotes, as seen in the figure.
Conclusion
Based on this review of studies, the authors postulated that EVE-derived piRNA-guided gene silencing may represent a previously unknown innate immunity against viruses in eukaryotes, in a mechanism parallel to CRISPR-Cas in prokaryotes. However, there is limited experimental evidence which directly shows this mechanism, and thus studies in this field are a high priority. Considering the popularity of gene editing in the current scientific climate, the authors believe that it is necessary to further elucidate the consequences of viral/transposonal gene transfer and piRNA-mediated gene editing.
Glossary
- Nucleotide
- a genomic unit that constitutes DNA and RNA.
- Bornavirus
- a virus that causes zoonotic diseases, known primarily to cause encephalitis in horses and sheep. Fragments of bornavirus genes integrated into the genome of many mammals, including humans, millions of years ago.
- Antisense
- complementary to specific DNA or RNA. Non-coding RNA
- Non-coding RNA
- An RNA that does not encode protein, and functions to regulate target DNA or RNA.
- Arbovirus
- general term for viruses that are transmitted via mosquitoes or ticks.
- Koala retroviruses
- viruses that causes leukemia and lymphoma in koalas that integrated into the koala genome relatively recently.
- Prokaryotes
- organisms that emerged at the very beginning of biological evolution, such as single-celled organisms like bacteria, which have no nucleus and barely exposed DNA within the cell.
- CRISPR/Cas
- a system consisting of a repetitive sequence within prokaryotic genome that encodes CRISPR RNA, which binds to the Cas enzyme to target foreign (including viral) RNA and silence them, making an adaptive antiviral immunity.
- Transposon
- genes that move within the genome (“the jumping gene”).
Journal information
- Title
- “piRNA-Guided CRISPR-like Immunity in Eukaryotes”
- DOI
- 10.1016/j.it.2019.09.003
- Authors
- Youdiil Ophinni1, Umberto Palatini2, Yoshitake Hayashi1, Nicholas F. Parrish3, 4
- Division of Molecular Medicine and Medical Genetics, Department of Pathology, Kobe University Graduate School of Medicine
- Department of Biology and Biotechnology, University of Pavia
- Genome Immunobiology RIKEN Hakubi Research Team, Cluster for Pioneering Research, RIKEN
- Center for Integrative Medical Sciences, RIKEN
- Journal
- Trends in Immunology